|
11/15/2023 0 Comments Application of southern blotting 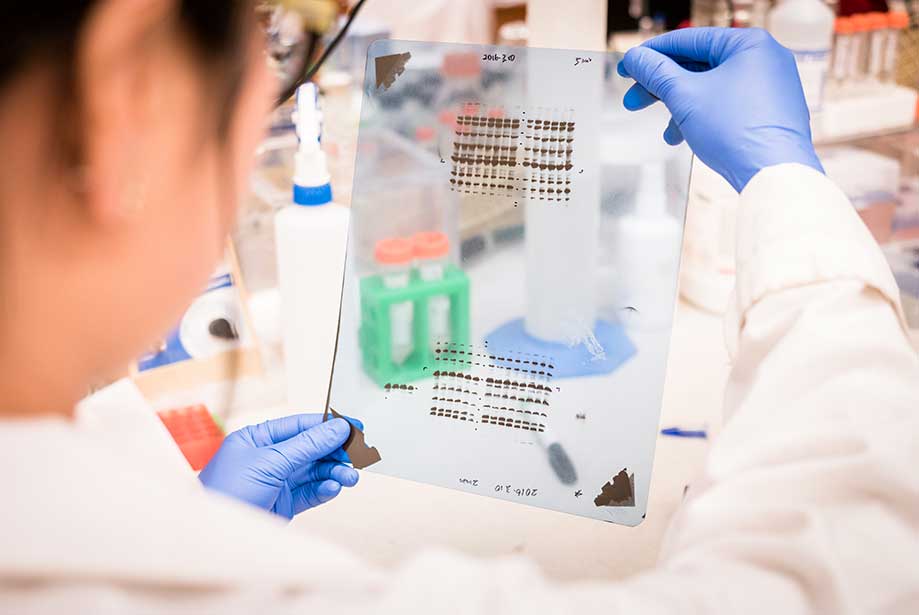 
The names for other blotting methods may follow this convention, by analogy. As the label is eponymous, Southern is capitalized, as is conventional of proper nouns. Other blotting methods (i.e., western blot, northern blot, eastern blot, southwestern blot) that employ similar principles, but using RNA or protein, have later been named for compass directions as a sort of pun from Southern's name. The method is named after the British biologist Edwin Southern, who first published it in 1975. The Southern blotting combines the transfer of electrophoresis-separated DNA fragments to a filter membrane in a process called blotting, and the subsequent fragment detection by probe hybridization. The tag allows any DNA fragments containing complementary sequences with the DNA probe sequence to be visualized within the Southern blot. The DNA fragments are transferred out of the gel or matrix onto a solid membrane, which is then exposed to a DNA probe labeled with a radioactive, fluorescent, or chemical tag. Briefly, purified DNA from a biological sample (such as blood or tissue) is digested with restriction enzymes, and the resulting DNA fragments are separated by using an electric current to move them through a sieve-like gel or matrix, which allows smaller fragments to move faster than larger fragments. This method is used in molecular biology. Southern blot is a method used for detection and quantification of a specific DNA sequence in DNA samples. Southern blot agarose gel under ultraviolet illumination. Southern blot membrane after hybridization and rinsing. Rapid Hybridization System-Multiprime, protocol booklet (1988) Amersham International plc.DNA analysis technique Agarose gel Tray with a stack consisting top down of a weight, paper towels, membrane of nitrocellulose or nylon, gel, salt solution and a slab of glass. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY. (1982) Molecular Cloning: A Laboratory Manual. (ed.) (1984) Methods in Molecular Biology, vol. 
J., eds.), IRL, Oxford and Washington DC, pp. (1985) Quantitative filter hybridization, in Nucleic Acid Hybridization: A Practical Approach (Hames, B. 9: Protocols in Human Molecular Genetics (Mathew, C. (1991) The detection of specific DNA sequences by enhanced chemiluminescence, in Methods in Molecular Biology, vol. (1984) Detection of mRNAs in sea urchin embryos by in situ hybridization using asymmetric RNA probes. (1984) Addendum: A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. (1983) A technique for radiolabelling DNA restriction endonuclease fragments to high specific activity. 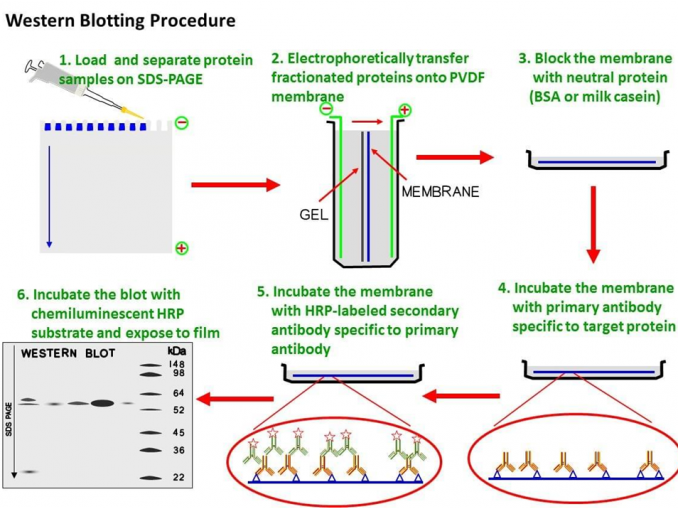
(1990) Filter hybridization and radiolabelling of nucleic acids, in Advances in Gene Technology vol. (1985) Rapid transfer of DNA from agarose gels to nylon membranes. (1984) Hybridization of nucleic acids immobilized on solid supports. (1988) Vacuum blotting enhances nucleic acid transfer. (1984) Detection of specific sequences-the Southern Transfer, in Methods in Molecular Biology, vol. (1991) Diagnosis of genetic disorders with linked DNA markers, in Methods in Molecular Biology, vol. (1991) DNA fingerprinting analysis: Methodology and its applications, in Methods in Molecular Biology, vol. (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
